Usage#
1. Data preparation#
Setup a demo directory in current working directory
import os demo_dir = os.getcwd() + "/SpecPipeDemo/"
Create a data directory and download real-world demo data
data_dir = demo_dir + "demo_data/" os.makedirs(data_dir) from swectral import download_demo_data download_demo_data(data_dir)
Create a directory for pipeline results
report_dir = demo_dir + "/demo_results_classification/" os.makedirs(report_dir)
2. Data configuration#
Create a SpecExp instance:
from swectral import SpecExp exp = SpecExp(report_dir)
The instance stores and organizes the data loading configurations of an experiment, which faciliates lazy-loading.
Check report directory:
exp.report_directory
Output:
'~/SpecPipeDemo/demo_results_classification/'
Add experiment groups:
exp.add_groups(['group_1', 'group_2'])
Add raster images:
exp.add_images_by_name(image_name="demo.", image_directory=data_dir, group="group_1") exp.add_images_by_name("demo.", data_dir, "group_2")
Output:
Following image items are added: Group Image Mask 0 group_1 demo.tiffLoad image ROIs using suffix to image names:
# By parameter name exp.add_rois_by_suffix(roi_filename_suffix="_[12].xml", search_directory=data_dir, group="group_1") # Or by parameter position exp.add_rois_by_suffix("_[345].xml", data_dir, "group_2")
Output:
Following ROI items loaded: Group Image ROI_name ROI_type ROI_source_file 0 group_1 demo.tiff 1-1 sample demo_1.xml 1 group_1 demo.tiff 1-2 sample demo_1.xml ... 9 group_1 demo.tiff 2-5 sample demo_2.xml
Show raster RGB preview with associated ROIs:
exp.show_image("demo.tiff", "group_1", rgb_band_index=(19, 12, 6), output_path=report_dir + "demo_rast_rgb1.png")
Output:
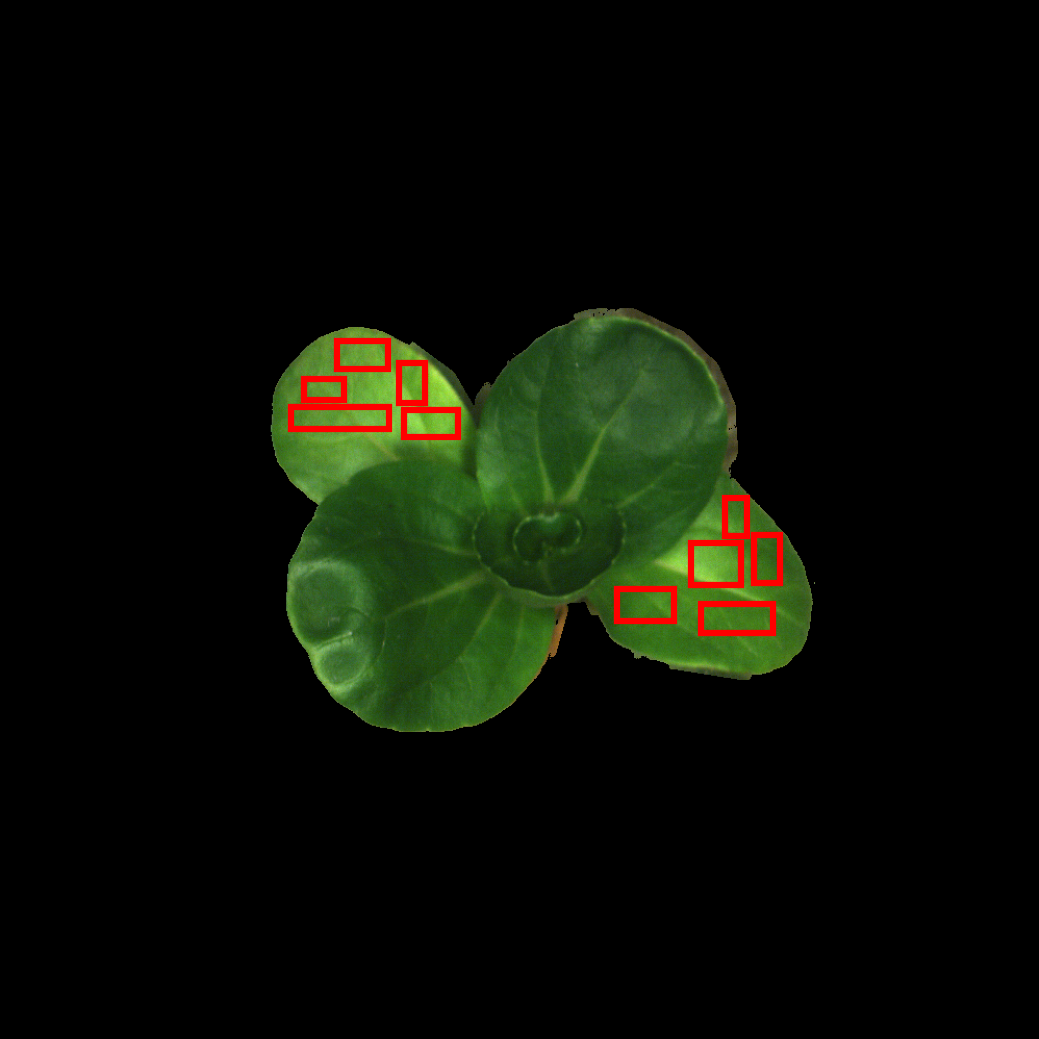

exp.show_image("demo.tiff", "group_2", rgb_band_index=(19, 12, 6), output_path=report_dir + "demo_rast_rgb2.png")
Output:
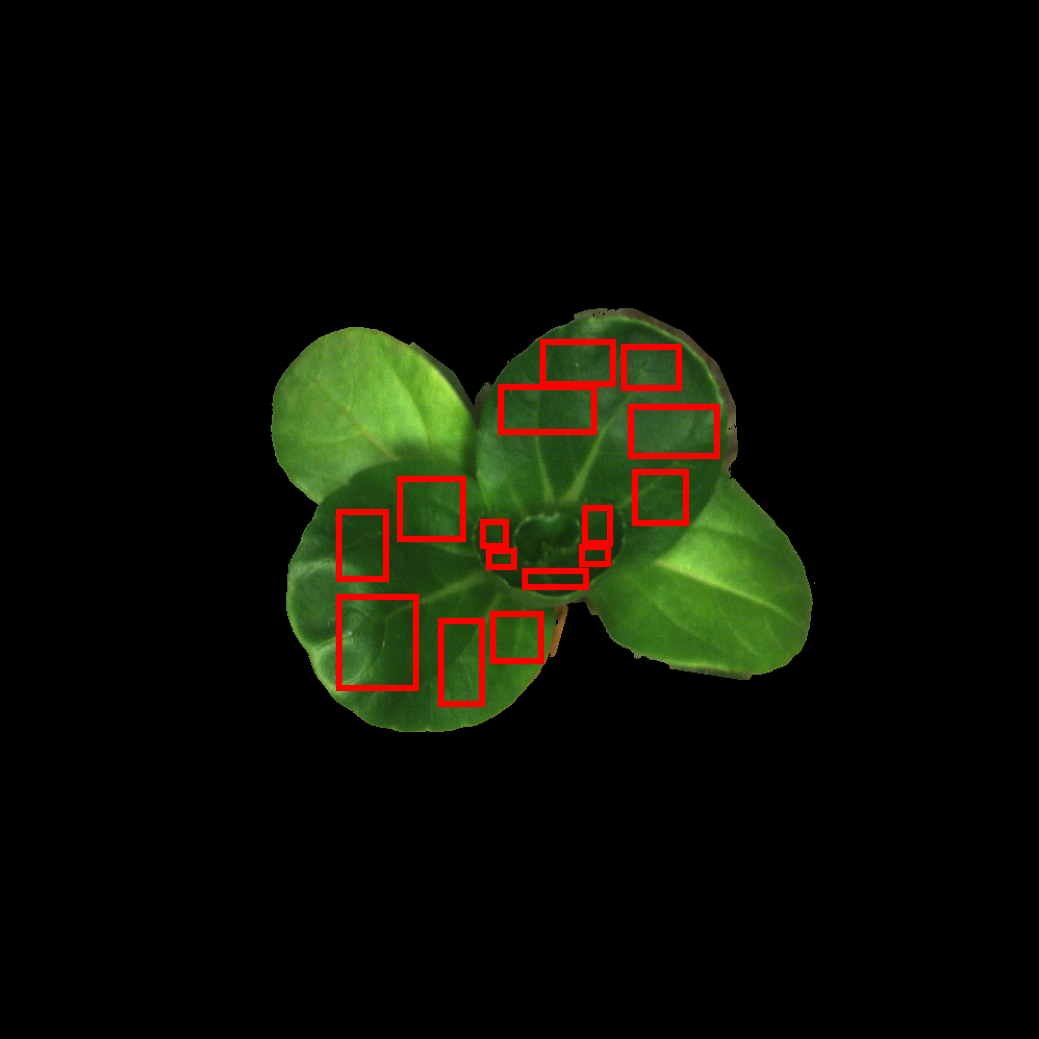

2.5. Sample labels and target values#
2.5.1 Set sample labels#
Get current sample label dataframe:
labels = exp.ls_labels()
Set new sample labels in the dataframe:
Here we use sample ROI names as sample labels:
labels.iloc[:, 1] = exp.ls_rois_sample(return_dataframe=True, print_result=False)["ROI_name"]
Update sample labels:
exp.sample_labels = labels
Check sample labels:
exp.ls_labels()["Label"]
Output:
0 1-1 1 1-2 ... 24 5-5
2.5.2 Set target values#
List target value dataframe:
targets = exp.ls_sample_targets()
Create mock target values for regression and update target dataframe:
Here we use leaf number:
targets["Target_value"] = [f"leaf_{labl[0]}" for labl in targets['Label']]
Load target values from updated target dataframe:
exp.sample_targets_from_df(targets)
Check target values:
exp.ls_targets()[["Label", "Target_value"]]
Output:
Label Target_value 0 1-1 leaf_1 1 1-2 leaf_1 ... 24 5-5 leaf_5
3. Design testing pipelines#
SpecPipe follows a structured data processing workflow with these sequential data levels:
Raster image data -> ROI spectra -> ROI statistics -> Traits to model
The data levels in SpecPipe includes:
Raster images: 0 - "image", input image path and output processed image path. 1 - "pixel_spec", if the process callable is applied to 1D spectrum of image pixel 2 - "pixel_specs_array", if the process callable is applied to 2D spectra array of image pixels 3 - "pixel_specs_tensor", if the process callable is applied to 3D spectra tensor of image pixels 4 - "pixel_hyperspecs_tensor", same as "pixel_specs_tensor" but optimized for hyperspectral images ROI spectra: 5 - "image_roi", raster with sample ROIs, for spectrum extraction 6 - "roispecs", 2D array of ROI spectra ROI statistics: 7 - "spec1d", arbitrary 1D data of samples, e.g. 1D spectra, flattened spectra statistical metrics Sample data: 8 - "assembly", sample data list for cross-sample interaction Models: 9 - "model", model evaluation with standard report output as filesThe corresponding data processing workflow is:
Raster image processing: 0 ~ 4 ↓ Extract ROI spectra: 5 - "image_roi" ↓ ROI spectra manipulation: 6 - "roispecs" ↓ Summarized ROI spectra: 7 - "spec1d" ↓ Sample assembly: 8 - "assembly" ↓ Modeling and model evaluation: 9 - "model"
The processing functions are incorporated in the pipeline according to the specified “data levels”. Parallel processes can be added with identical “data level” and “application sequence”, and they are arranged using full-factorial approach in the pipeline.
3.1 Create processing pipeline#
Create processing pipeline from SpecExp instance configured above:
from swectral import SpecPipe pipe = SpecPipe(exp)
3.2 Image processing#
Create some image processing functions, such as:
Standard normal variate:
from swectral.functions import snv
Pass-through method for comparison:
def raw(v): return v
3.3 ROI statistics#
Import spectral statistic metrics for ROI summary:
from swectral import roi_mean, roi_median
3.4 Add models to the pipeline#
Create some models:
from sklearn.ensemble import RandomForestClassifier from sklearn.neighbors import KNeighborsClassifier rf_classifier = RandomForestRegressor(n_estimators=10) knn_classifier = KNeighborsRegressor(n_neighbors=3)
3.5 Compose and check pipelines#
Compose pipelines:
pipe.build_pipeline( [ # 1 Image-wide baseline correction ((2, 2), [raw, snv]), # 2 ROI statistics ((5, 7), [roi_mean, roi_median]), # 3 Models (Feature selector included) ((7, 9), [rf_classifier, knn_classifier], {'validation_method': '2-fold'}) ] )
Check all processes including models:
pipe.ls_process()
Output:
Step_0 Step_1 Step_2 0 snv roi_mean KNeighborsClassifier 1 snv roi_mean RandomForestClassifier 2 snv roi_median KNeighborsClassifier 3 snv roi_median RandomForestClassifier 4 raw roi_mean KNeighborsClassifier 5 raw roi_mean RandomForestClassifier 6 raw roi_median KNeighborsClassifier 7 raw roi_median RandomForestClassifier
4 Execute pipelines#
Run:
pipe.run()
5 Generated reports#
Pipeline execution data is saved to local storage, use the methods to retrieve reports in the console:
result_summary = pipe.report_summary() chain_results = pipe.report_chains()
Check summary reports
The summary reports include:
result_summary.keys()
Output:
dict_keys([ 'Macro_avg_performance_summary', 'Marginal_macro_avg_AUC_stats_step_0', 'Marginal_macro_avg_AUC_stats_step_1', 'Marginal_macro_avg_AUC_stats_step_2', 'Marginal_micro_avg_AUC_stats_step_0', 'Marginal_micro_avg_AUC_stats_step_1', 'Marginal_micro_avg_AUC_stats_step_2', 'Micro_avg_performance_summary', 'sample_targets_stats'])Demonstration of macro-average performance metrics of classification:
result_summary['Macro_avg_performance_summary']
Output:
Step_0 Step_1 Step_2 Precision Recall F1_Score Accuracy AUC 0 2_0_%#1 5_0_%#1 7_0_%#1 0.86 0.84 0.84 0.94 0.95 ... 7 2_0_%#2 5_0_%#2 7_0_%#2 0.77 0.72 0.68 0.89 0.83
Demonstration of marginal macro-average performance metrics of classification:
result_summary['Marginal_macro_avg_AUC_stats_step_0']
Output:
Process_ID All 2_0_%#1 2_0_%#2 0 Process_label All snv raw 1 n_records 8 4 4 2 Mean_AUC_macro 0.85 0.95 0.76 3 Min_AUC_macro 0.63 0.94 0.63 4 Median_AUC_macro 0.91 0.95 0.76 5 Max_AUC_macro 0.97 0.97 0.87 6 p_vs_All 1.00 0.20 0.20 7 p_vs_raw 0.20 1.00 0.03 8 p_vs_snv 0.20 0.03 1.00 9 effect_vs_All 0.00 0.46 0.46 10 effect_vs_raw 0.46 0.00 0.94 11 effect_vs_snv 0.46 0.94 0.00
The processes of the step (here raw image and standard normal variates) are compared using non-parametric Wilcoxon signed-rank test.
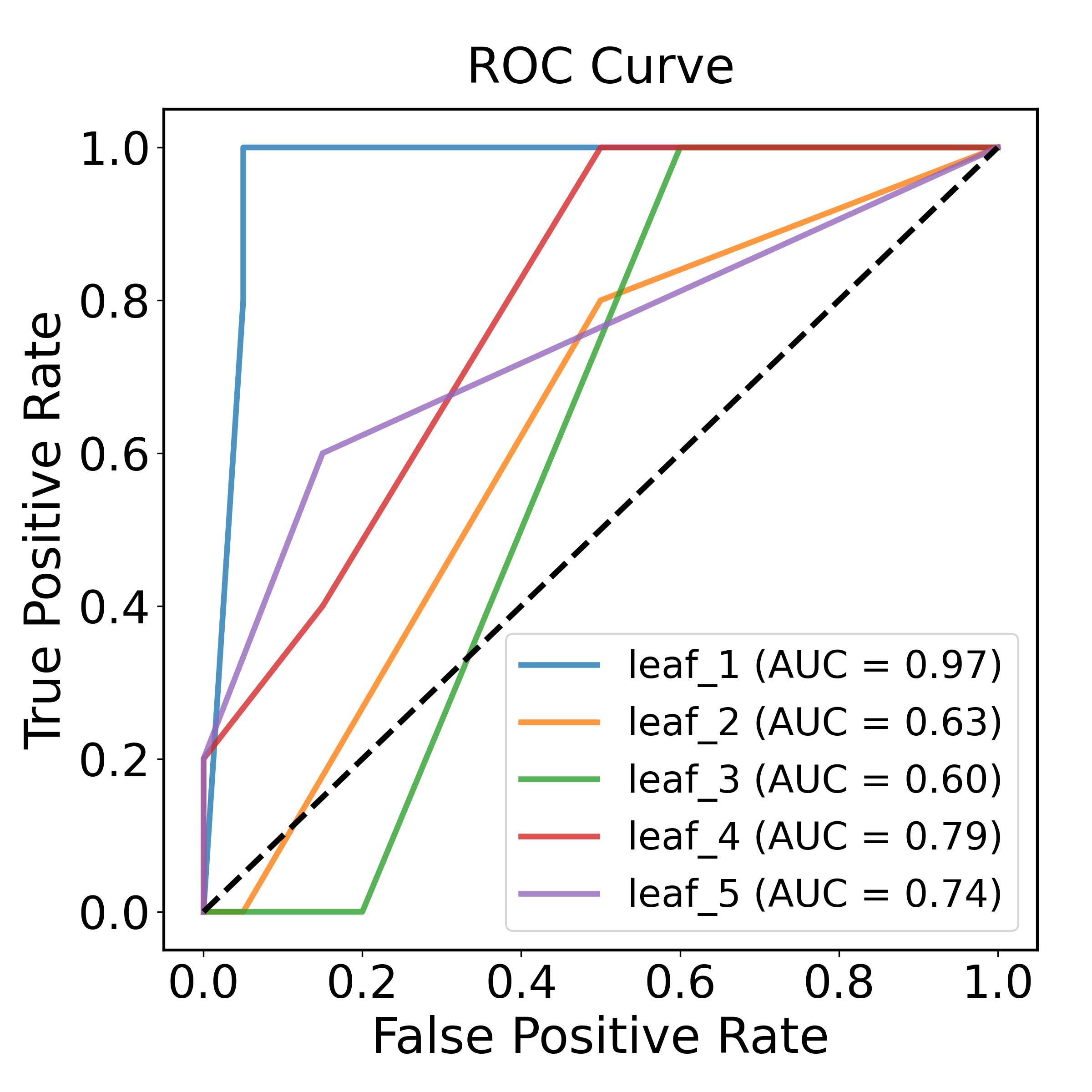
Demonstration of Receiver-Operating-Characteristic curve:
chain_results[0]['ROC_curve']
Output:
6 Regression demonstration#
6.1 Create a directory for regression results#
Create a directory for regression results
report_dir_reg = demo_dir + "/demo_results_regression/" os.makedirs(report_dir_reg)
6.2 Copy and update the previous pipelines to regression#
Copy and update SpecExp and SpecPipe instances
import copy exp_reg = copy.deepcopy(exp) pipe_reg = copy.deepcopy(pipe) targets_reg = copy.deepcopy(targets)
Update report directory of SpecExp
exp_reg.report_directory = report_dir_reg
Modify targets to numeric, here the numbers approaximate the age of the leaves
targets_reg["Target_value"] = [(5 - int(labl[0])) for labl in targets['Label']]
Specify the ROIs within a same leaf to a validation group to prevent data leakage
targets_reg["Validation_group"] = [f"leaf_{labl[0]}" for labl in targets['Label']]
Update target information using the modified target dataframe
exp_reg.sample_targets_from_df(targets_reg)
Check target values and validation groups
exp_reg.ls_targets()[["Label", "Target_value", "Validation_group"]]
6.3 Update the pipeline models to regressors#
Check and remove classification models
pipe_reg.ls_model() pipe_reg.rm_model()
Update the data manager
pipe_reg.spec_exp = exp_reg
Add regressors to the pipeline
Add some regressors:
from sklearn.ensemble import RandomForestRegressor from sklearn.neighbors import KNeighborsRegressor rf_regressor = RandomForestRegressor(n_estimators=10) knn_regressor = KNeighborsRegressor(n_neighbors=3) pipe_reg.add_model([knn_regressor, rf_regressor], validation_method="2-fold")
6.4 Execute regression pipelines#
Run:
pipe_reg.run()
6.5 Check results of regression pipelines#
Retrieve reports in console
result_summary_reg = pipe_reg.report_summary() chain_results_reg = pipe_reg.report_chains()
Check summary reports
The summary reports include:
result_summary_reg.keys()
Output:
dict_keys([ 'Marginal_R2_stats_step_0', 'Marginal_R2_stats_step_1', 'Marginal_R2_stats_step_2', 'Performance_summary', 'sample_targets_stats'])Demonstration of performance summary content:
result_summary_reg['Performance_summary'].columns
Output:
Index([ 'Step_0', 'Step_1', 'Step_2', 'Mean_Error', 'Standard_Deviation_of_Error', 'Mean_Absolute_Error', 'Normalized_MAE', 'CV_MAE', 'Mean_Squared_Error', 'Root_Mean_Squared_Error', 'Normalized_RMSE', 'CV_RMSE', 'Residual_Prediction_Deviation', 'R2' ], dtype='object')Check processing chain reports
For each chain, the reports include:
chain_results_reg[0].keys()
Output:
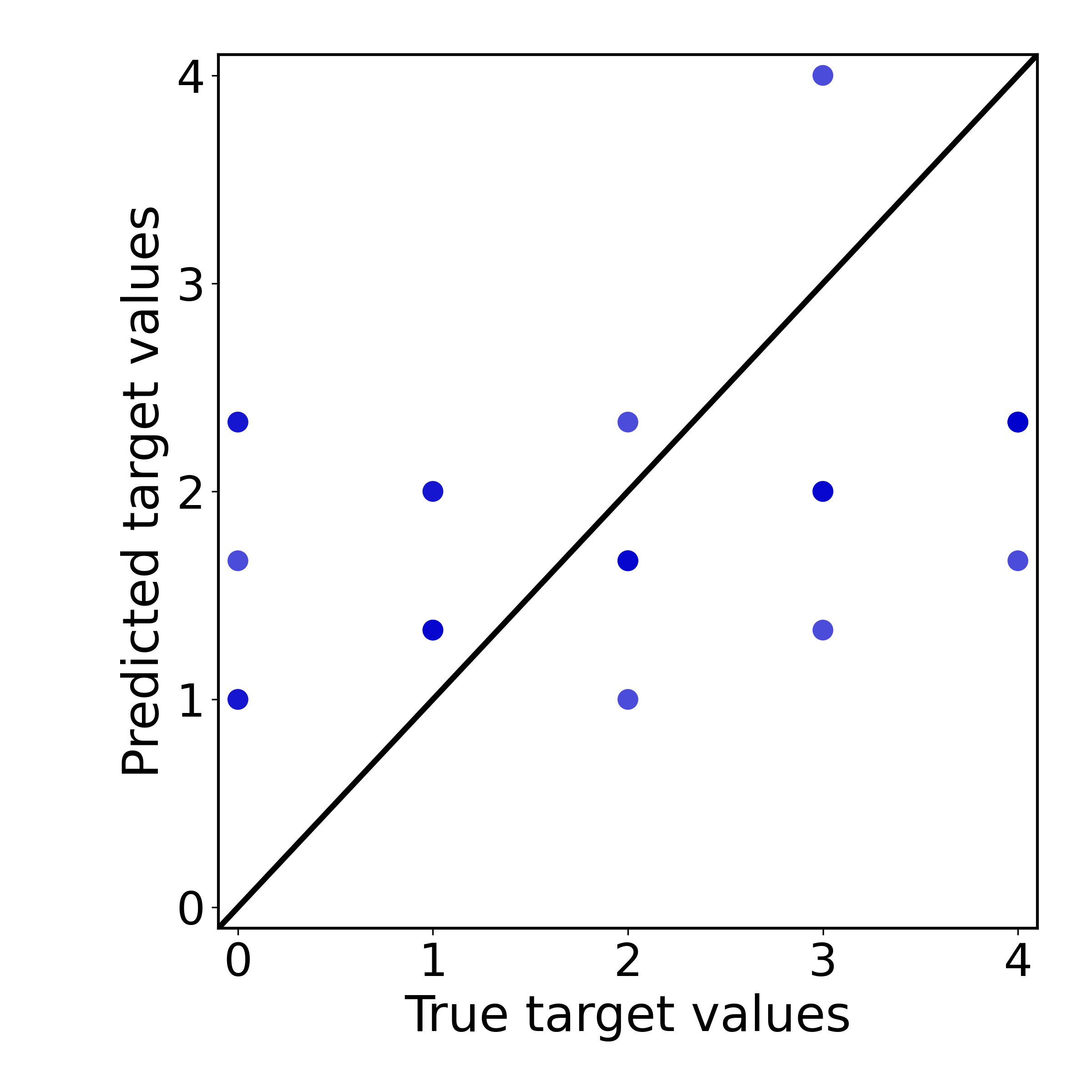
dict_keys([ 'Chain_processes', 'Regression_performance', 'Residual_analysis', 'Residual_plot', 'Scatter_plot', 'Validation_results'])Demonstration of the scatter plot of the processing chain:
chain_results_reg[0]['Scatter_plot']
Output:
7 Feature engineering fittable tests#
Feature engineering and resampling fittables (data transformers and resamplers) are fitted during the model validation process and function as integrated parts of the model. To incorporate these transformers, use the model connector functions combine_classifier or combine_regressor (similar to sklearn.pipeline.Pipeline but more flexible and enable swectral pipeline analysis).
This module includes a composer that generates batchwise combined models using a full factorial design:
from sklearn.preprocessing import StandardScaler from sklearn.feature_selection import SelectKBest, f_classif from swectral import IdentityTransformer # Passthrough transformer for comparison selector1 = SelectKBest(f_classif, k=5) # Select 5 features selector2 = IdentityTransformer() # For passthrough (no selection) from swectral import factorial_model_chains models = factorial_model_chains( [StandardScaler(), IdentityTransformer()], # Model step 1: test data scalers {'Feat5': selector1, 'FeatAll': selector2}, # Model step 2: test feature selection fittables # ... estimators={'KNN': knn_classifier, 'RF': rf_classifier}, # Estimators (specify custom labels using dictionary input) is_regression=False ) print(models)
Output:
[CombinedClassifier_StandardScaler_Feat5_KNN, CombinedClassifier_StandardScaler_Feat5_RF, CombinedClassifier_StandardScaler_FeatAll_KNN, CombinedClassifier_StandardScaler_FeatAll_RF, CombinedClassifier_IdentityTransformer_Feat5_KNN, CombinedClassifier_IdentityTransformer_Feat5_RF, CombinedClassifier_IdentityTransformer_FeatAll_KNN, CombinedClassifier_IdentityTransformer_FeatAll_RF]
Add the generated models to your pipeline:
pipe.add_model(models, validation_method="2-fold")
Tutorials / Demos#
You can try out SpecPipe using the following example scripts:
Basic Usage — demo_script_of_readme.py
Typical workflow — demo_script_typical_workflow.py
Parallel for Windows — demo_script_windows_parallel.py